Sommaire
- Influence of the hardness of bacteriological culture media on environmental monitoring with impaction type air samplers.
- A New Rapid Microbiology Method based on Measuring Oxygen. Depletion: An Assay for Testing Surfaces of Equipment, Facilities, and Personnel in Pharmaceutical Manufacturing Controlled Environments
- Cahier Pratique – Maîtrise de la qualité des gaz pharmaceutiques
- Sécurité microbiologique des produits cosmétiques : Enjeux et Réalités.
- Development of platform processes for the manufacture of Biopharmaceuticals.
- Développement d’un process Ultrafiltration / Diafiltration pour des applications de haute Concentration / Viscosité.
- Démarrage initial d’une salle propre et redémarrage après un évènement majeur.
The techniques available for the microbiological monitoring of products and manufacturing environments have come a long way in the last few decades. Even though enumerating microbial cells by counting colonies on an agar filled Petri dish is still a common technique, a variety of more sophisticated and rapid methods are making their way into the quality control microbiology laboratory.

These Rapid Microbiological Methods (RMM) exploit chemical and physical methods developed to elucidate the structure of cell components and the functions of biomolecules. These methods are then applied to the detection, enumeration, and identification of microorganisms that may be present in pharmaceutical, food and beverage, water and other samples submitted to the quality control labs. Thus, the simple view of a bacterial cell as a tiny living entity so small as to be invisible to the naked eye, and countable only by either microscopy or culture, is changing to a much more complex picture encompassing the most intimate details of the cell structure and the biomolecules that form it.
In this paper we describe an oxygen, depletion-based method as a rapid microbiology assay to test surface microbial contamination in cleanrooms environments. Other applications of this technology include testing microbial bioburden in raw materials, excipients, drug products, pharmaceutical water, and environmental monitoring applications in the pharmaceutical industry. The system detects microbial contamination based on measurements of oxygen depletion upon time of incubation in a pharmacopoeia-recommended liquid broth, such as TSB. The results of this study show that this rapid microbiology technology is reliable, sensitive, and equivalent to standard compendia tests.
Requirements to Monitor in Pharmaceutical Manufacturing Environments
The following is a bulleted summary of regulatory requirements for monitoring microbial contamination in pharmaceutical manufacturing environments:
• A proper detection system for microbial and total air particulate in cleanrooms.
• The implementation of an adequate environmental plan to provide information about the state of control in the manufacturing facility
• Adequate data evaluation required on both a short and long-term basis.
• The environmental monitoring plan should provide a good understanding and control of Heating, Ventilation and Air Conditioning (HVAC), HEPA filters and differential pressure issues.
• They should provide information on the amount and types of microorganisms recovered in the facility.
• They should be used as a supplementary method for evaluating a facility change control.
Microbial monitoring requirements in air and surfaces are:
• Air
1. Total Particulates (Inert or Nonviables, plus Viables)
a. Particle Counting (0.5 μm in diameter or larger)
2. Viables (typically bacterial and fungal cells and spores)
a. Settle or Settling Plates (Passive Air Sampling)
b. Air Samplers (Active Air Sampling)
• Surfaces (Viables)
1. Facilities (including walls, floors, etc.) and instruments
a. Contact Plates
b. Swabs
2. Personnel (Gloves, garment, etc.)
Microbial Surface Monitoring in Pharmaceutical Manufacturing Environments
Both the USA and the European Union GMPs require microbial surface monitoring in pharmaceutical cleanrooms. Section 18 of the EU aseptic manufacturing GMP document (Eudralex volume 4, Annex 1, 2008) establishes the following:
• Where aseptic operations are performed, frequent monitoring using methods such as settle plates, volumetric air and surface sampling (e.g. swabs and contact plates).
• Surfaces and personnel should be monitored after critical operations.
• Additional microbiological monitoring is also required outside production operations, e.g. after validation of systems, cleaning and sanitization.
Similarly, the USA Federal Drug Administration (FDA) GMP guidelines (Chapter V, section C, Monitoring Program) establishes the following:
• Personnel can significantly affect the quality of the environment in which the sterile product is processed. A vigilant and responsive personnel monitoring program should be established.
• Monitoring should be accomplished by obtaining surface samples of each operator’s gloves on a daily basis, or in association with each lot.
• This sampling should be accompanied by an appropriate sampling frequency for other strategically selected locations of the gown.
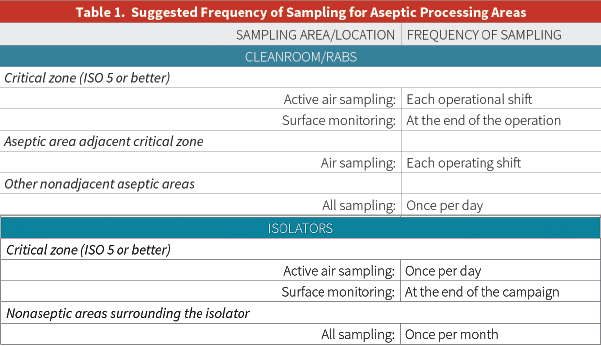
Moreover, the United States Pharmacopoeia (USP) alludes to microbial surface monitoring in its chapter <1116>, “Microbiological Control and Monitoring of Aseptic Processing Environments” (see Table 1 below):
A Rapid and Sensitive Microbiological Method Applied to Microbial Surface Monitoring
The SurCapt™ Microbial Surface Detection Kit is a new product from Particle Measuring Systems. This revolutionary technology uses swabs for microbial surface monitoring of:
• Surfaces
• Personnel garments
• Personnel gloves
• Equipment
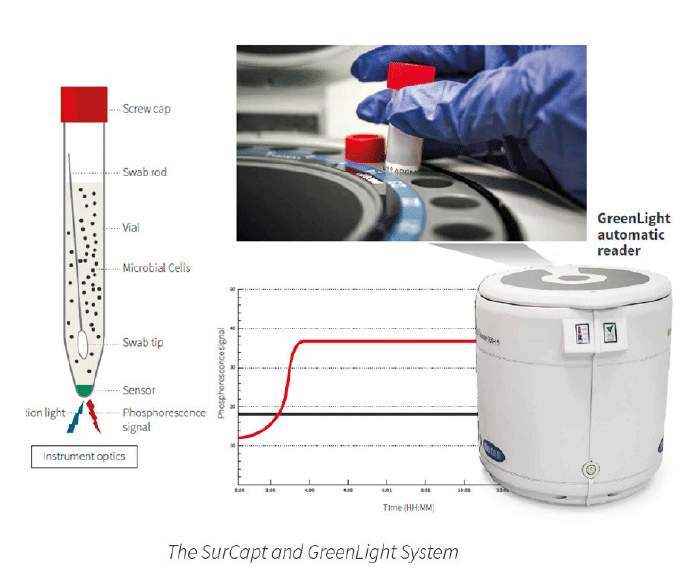
The SurCapt Kit uses GreenLight® Technology and detects microbial contamination based on measurements of oxygen depletion over incubation time in a pharmacopoeia recommended liquid broth, such as TSB.
The launch of this new rapid microbiological method coincides with the recent publication of the FDA draft guidance on the “Advancement of Emerging Technology Applications to Modernize the Pharmaceutical Manufacturing Base” (December 23, 2015). The guidance discusses a new FDA program that allows pharmaceutical companies to present proposals with a focus on innovative capability, such as a new testing tool, process, or proposed technology. The new guidance will facilitate the implementation of new microbiological methods in the pharmaceutical manufacturing arena.
Each SurCapt Microbial Surface Detection Kit is composed of the following:
1. 15 ml conical SurCapt vial with a sensor using GreenLight technology, filled with 10 ml of TSB.
2. Original Copan tube with a pre-moistening sponge embedded in 1 ml of SRK™ buffer solution and a FLOQyyySwab™.
3. PMS FLOQswab Kit.
The SRK buffer composition neutralizes the disinfectants present on the sampling surface and enhances the recovery and survival of microorganisms for approximately 18 hours.
The FLOQSwab included in each kit has an optimized recovery and release of 70-85%. This recovery is demonstrably higher than a number of other swabs and contact plates.
How Does Oxygen Sensing Work?
| The technology is based on the growth of the contaminating microbial culture. The technology follows this process: 1. Bacteria grow in a sample (incubated in a SurCapt vial) and consume dissolved O2. 2. The polymer sensor attached to the inside of the vial bottom reacts to the O2 depletion. 3. The measure of bacterial O2 consumption equates to microbial load. The greater the initial microbial load, the faster the result. |
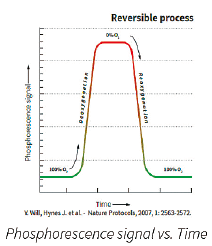
 | Basic Principles of O2 Sensing: • A fluorescent O2 sensitive probe is present at the bottom of each vial. • When O2 concentration increases, it causes a reversible decrease in the phosphorescence signal. • Inversely, when microbial respiration is active, the oxygen concentration in the media decreases and the sensor produces a larger phosphorescence signal. |
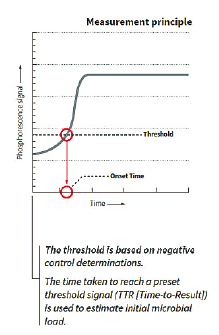
The graphical representation of the phosphorescence signal versus time (seen above) mimics a typical microbial growth curve. If the sample has a microbial contamination, the time that it takes the phosphorescent signal to cross a predetermined threshold is the Time-To-Result (TTR). The TTR for a large number of bacterial, yeast, and mold cultures tested is less than 24 hours.
Compared to other growth based rapid microbiological methods, the TTR for SurCapt is significantly faster. This is due to high sensitivity of oxygen sensing probes when compared to other RMM assay technologies.
Microbial Detection Using SurCapt and the GreenLight Reader
The internal SurCapt LED sends light that excites the sensor and the GreenLight reader’s optical system detects the fluorescence signal emission. This emission inversely correlates with the sample’s decrease in oxygen levels, giving results in microseconds.
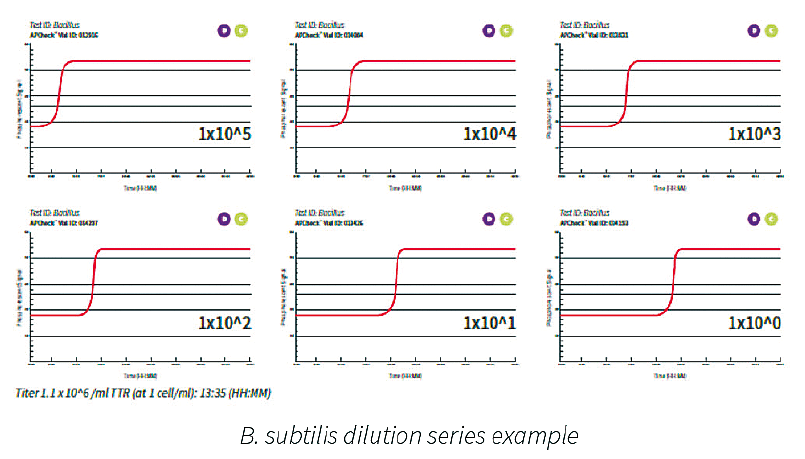
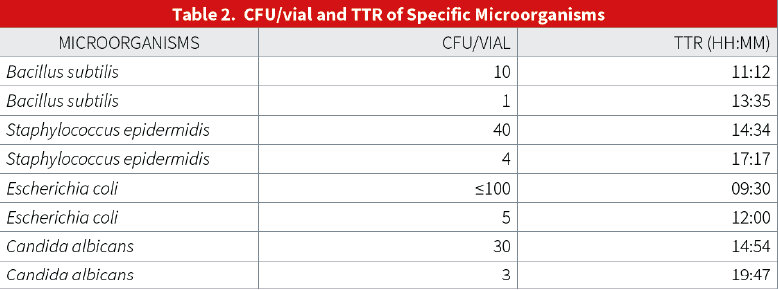
A typical dilution series of a microbial species demonstrated excellent linearity over a broad range of cell counts. An example for a B. subtilis dilution series is shown below. The range analyzed is 1x10E5 to 1x10E⁰ cells present in the test tube. The more bacterial cells present, the shorter time to result.
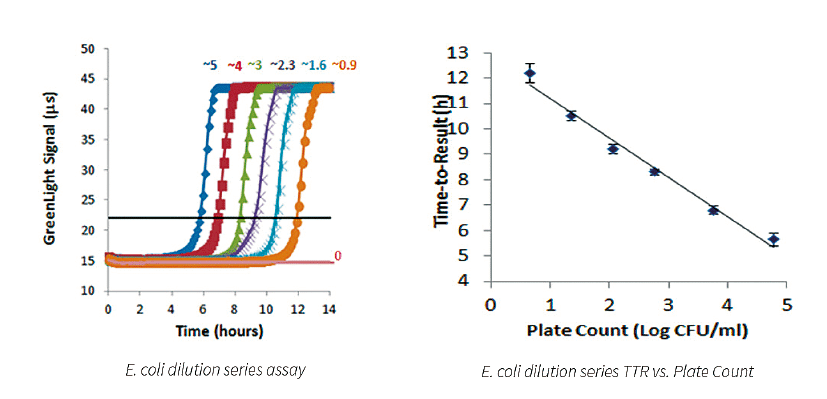
E. coli Dilution Series
The results shown above demonstrate that the assay has a tight standard deviation and good repeatability across a broad range of cell counts. There is an excellent correlation of TTR and plate count. Data suggests that the system is capable of detecting less than 5 CFU/ml with a TTR of approximately 12 hours. The results suggest that this system is significantly more sensitive (and faster) when compared to other growthbased rapid methods already on the market.
SurCapt (Specificity, Limit of Detection)
Preliminary validation results show that SurCapt has good sensitivity and LOD parameters. The TTR is always under 24 hours.
Conclusions
• The SurCapt Microbial Surface Detection Kit and GreenLight system represent a reliable and sensitive rapid microbial detection technology applied to pharmaceutical surface samples.
• The SurCapt Kit is comprised of high recovery flocked swabs, an oxygen-sensitive sensor and the traditional TSB liquid media for a ready-to-use disposable test (Refer to the PMSspecific GreenLight Operations Manual for further information).
• High recovery and optimal releasing of surface microbial contaminants using flocked swabs.
• Objective detection level results are as low as 1 cfu.
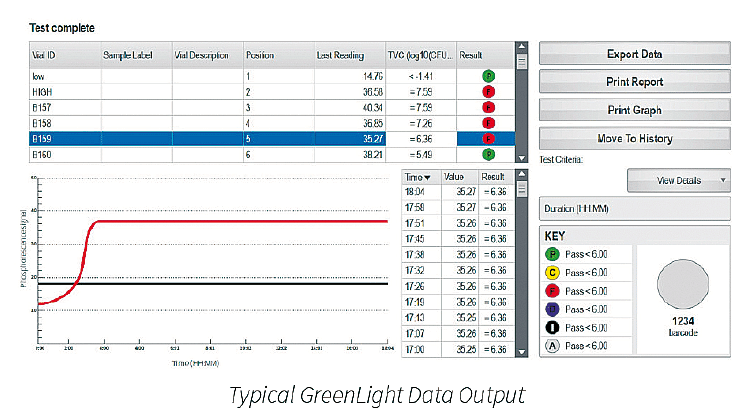
• The automatic reader eliminates human error in reading results and can sequentially operate up to six carousels, each carrying 24 surface samples.
• Improved traceability of vials and carousels with the GreenLight’s integrated barcode reader.
• The described method is non-destructive. The sample is available and intact for any further identification steps if required.
• The time taken to reach a preset threshold signal is faster than other methods, allowing a Presence/Absence result typically within 24 hours for surface samples from a Grade A / ISO 5 environment.

Claudio DENOYA – PARTICLE MEASURING SYSTEMS

Gilberto DALMASO – PARTICLE MEASURING SYSTEMS
Partager l’article
Références
• Patenteral Drug Association (PDA): www.pda.org
• Food and drug Administration (FDA): www.fda.gov
• United Stated Pharmacopoeia (USP): www.usp.org
• Particle Measuring Systems: www.pmeasuring.com
• European Pharmacopeia, EP 5.1.6 “Alternative Methods for Control of Microbiological Quality”
• PDA Journal of Pharmaceutical Science and Technology May/June 2008 vol. 62 no. 3 191-199
• USP chapter <1116> Microbial Monitoring in Aseptic Environments